The parameters of the current model can be saved or retrieved using the buttons illustrated below.
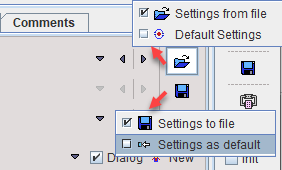
Model Default Configuration
Each model has a set of default parameter values. Initially, they are factory settings, but they can be re-defined by the user if he wants to establish a default configuration which is more adequate from him.
Settings as default |
Save the current parameter configuration as the new default for the particular model type. Included are the values, the fit flags, and the restrictions of all parameters |
Default Settings |
Resets the configuration of the current model to the saved defaults. |
Model Configuration File
A model configuration can also be saved as an explicit definition file (.kmModel). It includes the tissue model as well as all blood models used. Included are the initial parameters, the fit flags, the restrictions and the residual weighting scheme (except for the prescribed weighting with the explicit weighting factors).
Settings to file |
Save the current model and parameter configuration in a file. |
Default from file |
Retrieve a model configuration. Note, if the model in the file is different from the current model, the model is switched accordingly. |
Configuration files make it simple to establish defaults for particular data processing tasks: just load the appropriate .kmModel file and set it as the default. Another application is batch processing. There, multiple models can be fitted to the data by simply selecting the corresponding .kmModel files.